The risk for complex diseases such as cancer and other age-related pathologies varies across individuals and over time, arising from the intricate interplay of genetics and nongenetic factors. The epigenome records many of these interactions and shapes cellular fate, representing an attractive indicator of lifetime exposures, cellular state, and disease risk.
The Herzog group explores how the methylome can be harnessed to understand formation of age-related disease and improve disease risk prediction in non-invasive samples. The work is driven broadly by the following questions:
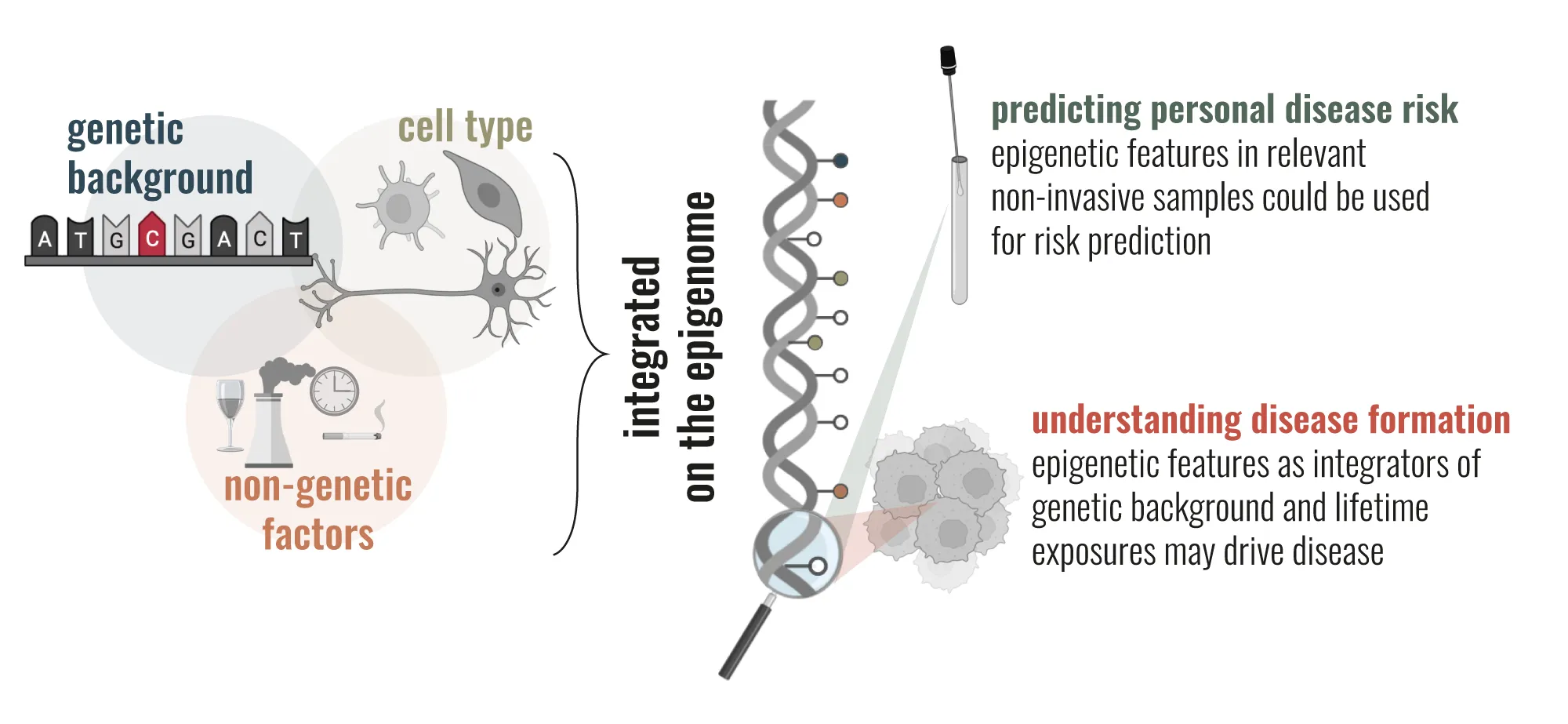
- How are genetic background, cell type, and exposures integrated and recorded in epigenetic ‘footprints’ that either directly predispose to, or are associated with, certain diseases?
- Can molecular ‘footprints’, in particular DNA methylation (DNAme) in non-invasively collected samples, detect current or future disease in other tissues of the same individual?
- What are the functional consequences of epigenetic alterations across diverse cell types, and how do they contribute to disease formation?
- Do different tissues within an individual age at different rates, and how is this associated with disease risk?
Using a combination of epigenetic and multi-omic data from human cohorts and laboratory research, we are tackling these aspects in a comprehensive and translational manner to ultimately better understand, predict, and prevent or treat age-related diseases to improve human health.
Group lead
Contact us
Department of Twin Research & Genetic Epidemiology,
King’s College London
St Thomas’ Hospital Campus
London SE1 7EH

